recent article in New Scientist titled ‘The Hidden Cause of Disease’ explores the idea that “The diseases most people die of have been attributed to unhealthy lifestyles. But evidence now suggests bacteria are to blame, heralding a revolution in medicine.”1 This is revolutionary thinking and for most people, too far out there. I understand that it may sound complicated or even simplistic, but that may be the beauty of it. The simplest explanation is usually the best and when the answer is too complex, the teacher is usually ‘culling the herd’ or really doesn't understand the concept that well themselves.
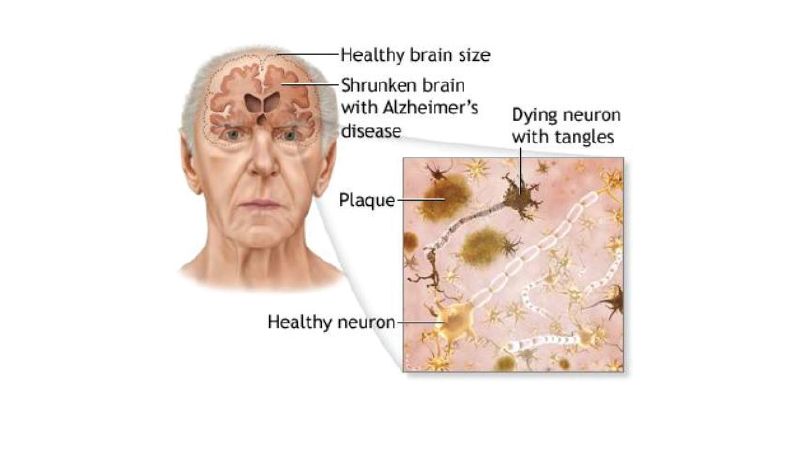
This can’t be truer than in the case of understanding the Human Microbiome. The current Human Microbiome Project, like the prior Human Genome Project, is revealing much information about us humans that has never been understood. “For decades, health experts have been lecturing us about our bad habits, blaming them for the surge in ‘lifestyle diseases’ like diabetes, stroke, and Alzheimer’s. Too much red meat, too little fruit and vegetables, smoking, drinking, obesity, and not enough exercise appear to make all these diseases more likely.
But no one really knows why, and we still haven’t worked out what causes any of them. Until now, bacteria’s involvement completely eluded us. But now DNA sequencing has revealed bacteria in places they were never supposed to be, manipulating inflammation in just the ways observed in these diseases. Some researchers, frustrated by years of failure to find causes, and therefore real treatments, for the diseases of ageing are cautiously excited. And with reason: this could change everything.
”1“ Our understanding of the link between the human microbiome and disease, including obesity, inflammatory bowel disease, arthritis, and autism, is rapidly expanding.
”2 These are quotes from just a couple of recent articles. Mind altering concepts, right? But what on earth does it have to do with wastewater and ‘Bacteria, Protozoa and Metazoa’. Good question. It is hard to find an accurate count of the number of Water Resource Recovery Facilities (WRRF) there are in this country, but it would seem to number around 16,000. (If you have a more accurate number I would love to hear it.) The EPA published this about the volume of water and it’s a lot. “Wastewater treatment facilities in the United States process approximately 34 billion gallons of wastewater every day. Wastewater contains nitrogen and phosphorus from human waste, food and certain soaps and detergents.
” 3 My point is that, each stream, river, lake, wetland and WRRF, not only possesses their own ecosystem, but their own microbiome. As the technology has increased our own ability to understand this has increased as well. So we can perform genomic analysis of each human, each stream, each river, each lake, each wetland, and each WRRF. Also, may I say, each process within a facility? As we do this we may better understand the microbiome of each WRRF and why we don’t always get the same outcome when we seemingly follow the same procedure/process and get different results.
Within the limitations of the accuracy of most analytical methods that we are forced to follow, this might provide an explanation of why two WRRF can have similar waste streams with similar concentrations of BOD, TSS, and Toxicities and have completely different results when the effluent is analyzed. Same food, but a different cast of characters, largely the bacteria, are consuming and metabolizing the
food differently.
In the same way, the protozoa and metazoa are then feeding on bacteria that may be similar, but are actually different, based on nutritional value, genetic type, and the process you are using and testing from. Therefore, if we are looking for the cause of an illness in the process, the cause may be as simple as not having the right bacteria to treat a new waste stream. In the case of the nitrifiers, it may take weeks to months to develop sufficient populations to treatment your volume of sewage, but they will develop. The organisms are there; they just need to build a suitably large population. The current accepted technologies to test for these types of organisms can take weeks to months, as well. However, with genomics we can follow these changes in days.
Just one example. Our WRRF we are required to analyze five times a week for E. coli and a limit of 126, Average Monthly and 157, Average Weekly, based on No./100mL. This is still a difficult parameter for some facilities to meet. Using genomic testing, we found only one species of identifiable E. coli, but nearly 2100 total species of identifiable bacteria. Imagine understanding what that really means to the receiving body of water.
What does it all mean? Just that if you work out and you don’t get the same result as someone else there may be an explanation if we could understand your microbiome. Diabetes may be in your family, but why did you get it and your sister didn’t. The same goes for a WRRF. If you don’t get the same results at your facility as the same plant elsewhere gets, maybe understanding the treatment plants microbiome may help you out. In a future article, we will expand on this and more about the protozoan and metazoan connection.
1 www.newscientist.com, New Scientist Ltd, Registered Office: 25 Bedford Street, London, WC2E 9ES.
2 www.nature.com/articles/nm.4517
3 www.epa.gov/nutrientpollution/sources-and-solutions-wastewater